.jpg?1775424008)
2nd ISFMS - Non-Coding RNAs and Epigenetics in Cancer
Part of the International Symposium on Frontiers in Molecular Science series
21–23 June 2017, Basel, Switzerland
noncoding RNA, Epigenetics, Cancer
- Go to the Sessions
- Event Details
-
- Conference Chairs
- Sessions
- Instructions for Authors
- Sponsors and Partners
- Conference Venue
- Accomodation
- Welcome Message by Regierungsrat Dr. Conradin Cramer
- List of Participants
- Conference Photos
- Conference Award
- Conference Video
- Travel & Registration Information
- Conference Organizers
- Conference Schedule
- Events in series ISFMS
Conference Chairs
gcalin@mdanderson.org
burkhalter@mdpi.com
[Not defined]
degen@mdpi.com
Instructions for Authors
To register for this conference, please follow this link. If you need assistance, please email the Conference secretariat.
- Create an account on Sciforum.net and follow the procedure to "Submit a New Abstract" in your User Home.
- Indicate which thematic area is best suited for your poster.
- Submit a 200 word abstract in English.
Each presenter will be provided with an A0 vertical poster board (841 x 1189 mm / 33.1 x 46.8 in). Please print your poster prior to the conference.
A plan of the poster session will be circulated later on. Posters should be set up the first day of the conference (21 June) between 7:30 am and 11:00 am and remain up until 23 June – 11:00 am. The same day, posters must be removed by 1:40 pm, the posters remaining after this time will be removed and recycled.
- Special issue: "Selected Papers from the 2nd International Symposium on Frontiers in Molecular Science: Non-Coding RNAs and Epigenetics in Cancer" edited by Prof. Dr. Thomas Schmittgen in the journal IJMS.
IJMS is indexed by the Science Citation Index Expanded (Web of Science), MEDLINE (PubMed) and other databases and has an Impact Factor of 3.257 (2015). - All Panel abstracts, paper abstracts, posters and presentations will be available online in Open Access on Sciforum.net.
Noncoding RNAs and epigenetics continue to play fundamental roles in the etiology, detection and treatment of cancer. Over the past decade and a half, researchers have unravelled important roles for miRNA in cancer. Translational applications such as miRNA diagnostics have found their way to the clinic. More recently, a focus on lncRNAs and other noncoding RNAs has emerged including the ability of lncRNAs to bind to and sequester miRNAs by working as competing endogenous RNAs. Several anticancer drugs that inhibit DNA methylation and acetylation are under clinical evaluation. This Special Issue of International Journal of Molecular Sciences (IJMS) will highlight work presented at the Second International Symposium on Frontiers in Molecular Science “Non-Coding RNAs and Epigenetics in Cancer”. We invite all attendees to submit a full-length research article from their presented work or a review article of their choosing on the subject of epigenetics in cancer. Submitted work will be due early Fall 2017 with an anticipated publication date of December, 2017.
Special Issues Open for SubmissionRegulation by Non-Coding RNAs
Collection Editor: Martin Pichler
31 December 2017
Targeting Cancer through RNA Biology
Editor: Michael Ladomery
31 December 2017
DNA Methylation
Editor: Reinhard Dammann
31 December 2017
Tumor Microenvironment
Editor: Naofumi Mukaida
30 November 2017
Advances in Molecular Oncology
Collection Editor: William Chi-shing Cho
31 December 2017
Hormones-Dependent Cancers: New Aspects on Biochemistry and Molecular Pathology
Editor: Dr. Yasuhiro Miki
31 December 2017
Galectins in Cancer and Translational Medicine
Editor: Prof. Dr. Armando Bartolazzi
30 November 2017
Role of Telomeres and Telomerase in Cancer and Aging
Editor: Dr. Gabriele Saretzki
30 September 2017
hCG°™ An Endocrine, Regulator of Gestation and Caner
Editor: Prof. Dr. Udo Jeschke
30 September 2017
TGF-beta Family in Fibrosis and Cancer
Editor: Dr. Isabel Fabregat
30 September 2017
Cancer Stem Cells
Editor: Dr. Hiroyuki Tomita, Prof. Dr. Masahito Shimizu, Prof. Dr. Takuji Tanaka
30 June 2017
Conference Venue
Conference Venue
The conference will be held at the BioCenter of the University of Basel. The address is Klingelbergstrasse 70, Basel.
Basel
Basel is the third largest city in Switzerland and is located in the Northeast of the country, on the border of France and Germany. It is an important economic center in Switzerland and offers a rich cultural life. It is home to the Museum of Fine Arts, which holds the most significant and largest collection of art in Switzerland, the Foundation Beyeler, the Theater Basel and the Musical Theater Basel.
Located on the bank of the Rhine and situated only a few kilometers from the German Black Forest and French Vosges, it also offers great possibilities for excursions in the nature.
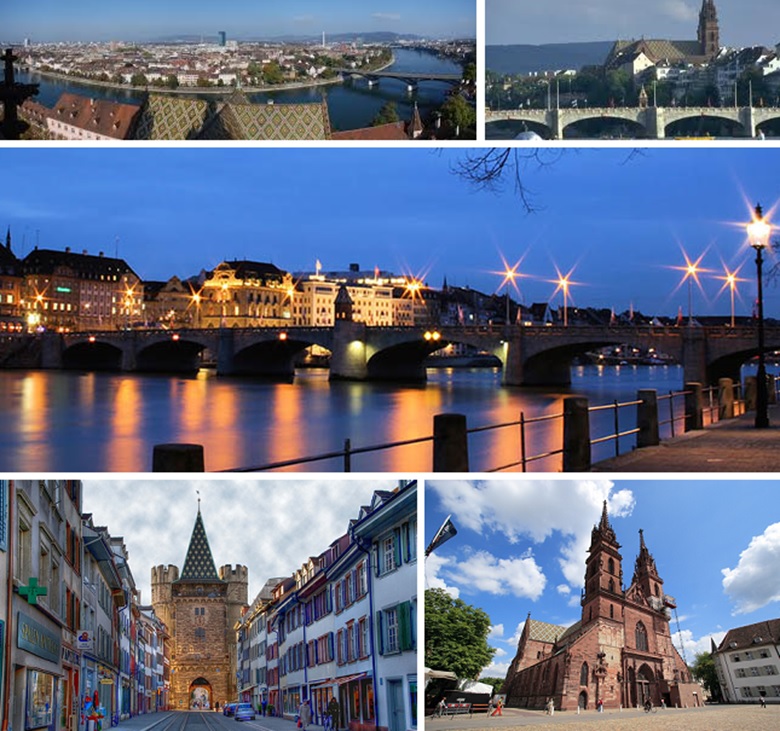

University of Basel
The University of Basel was founded in 1460 and is the oldest university in Switzerland. It prides itself in its tradition of over five hundred years of excellence in teaching, learning, and research, during which it has adopted a forward-looking approach to new developments in science to provide high-quality education and underpin its well-deserved reputation as a university capable of attracting staff and students from all over the world. Located in the heart of Basel, and drawing on innovation anchored in tradition, the University of Basel is dynamic and innovative. Research at the University of Basel focuses on the life sciences and culture. Both core areas comprise a wide range of projects, involving research staff and scientists committed to interdisciplinary work and a cross-faculty approach to advancing the knowledge of life and the workings of culture. The University of Basel is committed to providing state-of-the-art education and to facilitating outstanding research. (Adapted from www.topuniversities.com).

Accomodation
Basel Tourismus
Basel Tourismus kindly offers participants of the ISFMS2017 Conference the possibility to use their booking system to obtain the best rates for Hotels in Basel. Click the link to the booking system to reserve a room.
Welcome Message by Regierungsrat Dr. Conradin Cramer
Welcome to Basel! We are proud to host such an eminent group of scientists in our city. On the other hand, we may modestly say, that this location is ideal for a high level symposium on molecular science in general and on cancer medication in particular.
Basel is a hot spot for Life Sciences. The Life Sciences form the basis of our economy and are an important source of our public wealth. Moreover Basel has an all-embracing scientific ecosystem in the Life Sciences. The two global players Novartis and Roche alone contribute a volume of about 6 billion Swiss francs from Life Science Research. Although financed at a much lower level our University has a strong focus and hits well above its size in the Life Sciences – a knowledge pipeline beginning with the basic research at the Biozentrum and continuing along a translational path to the clinical research and practical application in the University Hospitals. The University is associated with renowned research institutes such as the Friedrich Miescher Institute of Novartis and the Swiss Tropical and Public Health Institute (Swiss TPH), which, like the University is a public institution. Of course the University also cooperates with top Universities in Switzerland – especially the ETH Zurich – and worldwide. There is also a close cooperation with our regional University of Applied Sciences.
If you look out of the window you will see the Life Science Campus arising, on which you soon will find the ETH-Department of Bio Systems and Systems Engineering in close vicinity to our university-institutes, such as the departments of Chemistry and Physics, the Biozentrum and the Department of Biomedical Science. In the same area we find the campus of our University hospital. We thus combine all of the essential components for a successful translation from high end research to clinical application in an area less than a square kilometre.
How can this be done by a little administrative region such as Basel City with only 190‘000 inhabitants? It is possible because of strong regional partnerships, especially with our neighboring county, that sends the greatest cohort of students to the University of Basel. Our entire region is proud of its university and participates to its wellbeing, financially, culturally and emotionally. And of course there are the subsidies of the federal government as well as national and international research grants which are acquired with great success by our university and provide the fuel for the research engine. The whole ecosystem is embedded in a trinational region, where the three nations France, Germany and Switzerland meet and form one of the strongest Life-Science clusters of the world. A region of frontiers, ideally prepared to test and move beyond the frontiers in molecular sciences!
In Basel you encounter a curious combination of small town with a strong identity and a worldwide scientific impact. We believe this to be a unique strength of this area. The short distance between centres of excellence facilitates free exchange between scientists and leads to an openness to other cultures and a profound commitment to the improvement of life quality worldwide. Your meeting here is a fine signal of this Ambition.
With this brief welcome note I hope to have shown, that you have chosen excellent surroundings for your congress. I wish you inspiration, a good atmosphere and of course scientific success. However, please don’t forget to explore our other highlights: the fine architecture of Basel, the cultural highlights and, of course, the culinary opportunities. In short: I hope you will also find a little time for leisure and enjoyment during your stay here.
List of Participants
Aeschimann Florian
Friedrich Miescher Institute Basel, Switzerland
florian.aeschimann@fmi.ch
Agami Reuven
Netherlands Cancer Institute, Amsterdam, The Netherlands
r.agami@nki.nl
Angermaier Johanna
Charité Universitätsmedizin Berlin, Germany
johanna.angermaier@charite.de
Askarian-Amiri Marjan
University of Auckland, New Zealand
m.askarian-amiri@auckland.ac.nz
Avni Dror
Sheba Medical Center, Tel Hashomer, Israel
droravni@msn.com
Bar-Eli Menashe
The University of Texas, Houston, USA
mbareli@mdanderson.org
Battino Maurizio
Università Politecnica delle Marche, Italy
m.a.battino@univpm.it
Behera Alok
ETH Zurich, Switzerland
alok.behera@pharma.ethz.ch
Berger Stefanie
University of Basel, Switzerland
st.berger@unibas.ch
Biswas Roopa
Walter Reed National Military Medical Center, Bethesda, USA
roopa.biswas@usuhs.edu
Bjørklund Sunniva
Oslo University Hospital, Oslo, Norway
sunniva.bjorklund@gmail.com
Boeri Mattia
Fondazione IRCCS Istituto Nazionale dei Tumori, Milan, Italy
mattia.boeri@istitutotumori.mi.it
Bojar Daniel
ETH Zurich, Switzerland
daniel.bojar@bsse.ethz.ch
Brancati Giovanna
Friedrich Miescher Institute Basel, Switzerland
giovanna.brancati@fmi.ch
Burkhalter Matthias
MDPI Basel, Switzerland
burkhalter@mdpi.com
Bustin Stephen
Anglia Ruskin University, UK
stephen.bustin@anglia.ac.uk
Calin George A.
The University of Texas, Houston, USA
gcalin@mdanderson.org
Candi Eleonora
University of Rome, Rome, Italy
candi@uniroma2.it
Cascione Luciano
Institute of Oncology Research, Bellinzona, Switzerland
luciano.cascione@ior.iosi.ch
Catuogno Silvia
Istituto per l'endocrinologia e l'oncologia, Napoli, Italy
silviacatuogno@virgilio.it
Chen Yue
MDPI Wuhan, China
yue.chen@mdpi.com
Coe Elizabeth
University of Bath, UK
eac51@bath.ac.uk
Coscujuela Tarrero Lucia
University of Turin, Orbassano, Italy
lucykosky@gmail.com
Cramer Conradin
Regierungsrat Kanton Basel-Stadt
conradin.cramer@bs.ch
Cristina Vinci
University of Milan, Italy
cristina.vinci@unimi.it
Croce Carlo
The Ohio State University, Columbus, USA
carlo.croce@osumc.edu
Cuculovic Milos
MDPI Basel, Switzerland
cuculovic@mdpi.com
Daum Janine
Friedrich Miescher Institute Basel, Switzerland
janine.daum@fmi.ch
Degen Michèle
MDPI Basel, Switzerland
degen@mdpi.com
Diederichs Sven
Deutsches Krebsforschungszentrum, Heidelberg, Germany
s.diederichs@dkfz.de
Eichmüller Stefan
Deutsches Krebsforschungszentrum, Heidelberg, Germany
s.eichmueller@dkfz.de
Enguita Francisco
University of Lisbon, Portugal
fenguita@fm.ul.pt
Esteller Manel
Bellvitge Biomedical Research Institute, Barcelona, Spain
mesteller@idibell.cat
Faridani Omid
Karolinska Institutet, Sweden
omid.faridani@licr.ki.se
Felley-Bosco Emanuela
University of Zurich, Switzerlandemanuela.felley-bosco@usz.ch
Ferracin Manuela
University of Bologna, Bologna, Italy
manuela.ferracin@unibo.it
Ferrero Giulio
University of Turin, Torino, Itay
giulio.ferrero@unito.it
Fodor Barna
Novartis Institutes for BioMedical Research, Basel, Switzerland
barna.fodor@novartis.com
Freeland Alistair
MDPI Basel, Switzerland
freeland@mdpi.com
Fuentes-Mattei Enrique
MD Texas Cancer Center, Houston, USA
efuentes1@mdanderson.org
Ghoshal Kalpana
The Ohio State University, Columbus, USA
ghoshal.1@osu.edu
Grosshans Helge
Friedrich Miescher Institute for Biomedical Research, Basel, Switzerland
helge.grosshans@fmi.ch
Gruber Andreas
University of Basel, Basel, Switzerland
aj.gruber@unibas.ch
Guerin Delphine
MDPI Basel, Switzerland
guerin@mdpi.com
Haidar Ziyad
University of los Andes, Santiago de Chile, Chile
zhaidar78@gmail.com
Hauer Christian
Novartis, Basel, Switzerland
christian.hauer@novartis.com
Hellio Claire
Université de Bretagne Occidentale, France
claire.hellio@univ-brest.fr
Huang Qihong
The Wistar Institute, Philadelphia, USA
qhuang@Wistar.org
Hwang Dae-Yong
Konkuk University Medical Center, Seoul, Korea
hwangcrc@kuh.ac.kr
Jeronimo Carmen
University of Porto, Porto, Portugal
carmenjeronimo@ipoporto.min-saude.pt
Jia Xiaofeng
University of Maryland, USA
XJia@som.umaryland.edu
Jiang Yanrui
ETH Zurich, Switzerland
yanrui.jiang@bsse.ethz.ch
Johnson Rory
University of Bern, Switzerland
rory.johnson@dkf.unibe.ch
Kancherla Venkatesh
Universitätsspital Basel, Switzerland
venky198922@gmail.com
Klajic Jovana
Akershus University Hospital, Denmark
jjovana38@yahoo.com
Kluiver Joost
University Medical Center Groningen, Groningen, The Netherlands
j.l.kluiver@umcg.nl
Konert Madlen
GATC Biotech, Konstanz, Germany
m.konert@gatc-biotech.com
Kresoja-Rakic Jelena
University of Zurich, Switzerland
jelena.rakic@usz.ch
Krummheuer Jörg
Exiqon A/S, Vedbæk, Denmark
jgk@exiqon.com
Kunej Tanja
University of Ljubljana, Slovenia
tanja.kunej@bf.uni-lj.si
Kurek Roman
Exiqon
rok@exiqon.com
Li Qiao
University of Ottawa, Ottawa, Canada
qiaoli@uottawa.ca
Lin Sen
MDPI Basel, Switzerland
sen.lin@mdpi.com
Lin Shu-Kun
MDPI Basel, Switzerland
lin@mdpi.com
Lucic Matije
ETH Zurich, Zurich, Switzerland
matije.lucic@pharma.ethz.ch
Mapelli Sarah
Institute of Oncology Research, Bellinzona, Switzerland
sarah.mapelli@ior.iosi.ch
Martinez Maite Huarte
University of Navarra, Pamplona, Spain
clopezg@unav.es
Mattick John S.
University of New South Wales, Sydney, Australia
j.mattick@garvan.org.au
Mercado Nicolas
Novartis Institutes for BioMedical Research, Basel, Switzerland
nicolas.mercado@novartis.com
Mihaila Delia
MDPI Basel, Switzerland
mihaila@mdpi.com
Mowla Seyed Javad
Tarbiat Modares University, Tehran, Iran
sjmowla@modares.ac.ir
Muller Fabbri
University of Southern California, Los Angeles, USA
mfabbri@chla.usc.edu
Napoli Sara
Institute of Oncology Research, Bellinzona, Switzerland
sara.napoli@ior.iosi.ch
Nasrollahzadeh Sabet Mehrdad
University of Tehran, Tehran, Iran
dr.m.sabet@gmail.com
Negrini Massimo
University of Ferrara, Ferrara, Italy
ngm@unife.it
Noghero Alessio
University of Turin, Candiolo, Itay
alessio.noghero@ircc.it
Orom Ulf Andersson
Max Planck Institute for Molecular Genetics, Berlin, Germany
oerom@molgen.mpg.de
Pahlevan Kakhki Majid
Karolinska Institute, Solna, Sweden
Majid.pahlevan.kakhki@ki.se
Panatta Emanuele
University of Rome, Rome, Italy
panatta.emanuele@gmail.com
Parrella Paola
Laboratory of Oncology IRCCS Casa Sollievo della Sofferenza
pparrella@operapadrepio.it
Pasculli Barbara
Laboratory of Oncology IRCCS Casa Sollievo della Sofferenza
b.pasculli@operapadrepio.it
Pasquinelli Amy
University of California, San Diego, USA
apasquinelli@ucsd.edu
Pena Rodrigo
University of Zurich, Switzerland
rodrigo.pena@dmmd.uzh.ch
Pichler Martin
Medical University of Graz, Austria
martin.pichler@medunigraz.at
Porcellini Elisa
University of Bologna, Italy
elisa.porcellini3@unibo.it
Quintavalle Cristina
Universitätsspital Basel, Switzerland
cristina.quintavalle@usb.ch
Rani James Alva
Deutsches Krebsforschungszentrum, Heidelberg, Germany
alvarani@gmail.com
Ren Yong
MDPI Wuhan, China
yong.ren@mdpi.com
Reshadatian Marzieh
Ahvaz Jundishapur University of Medical Sciences, Ahvaz, Iran
m.reshadatian@yahoo.com
Rigoutsos Isidore
Thomas Jefferson University, Philadelphia, USA
Isidore.Rigoutsos@jefferson.edu
Rittman Martyn
MDPI Basel, Switzerland
rittman@mdpi.com
Russo Aniello
University of Campania Luigi Vanvitelli, Caserta, Italy
aniello.russo@unina2.it
Russo Lucia
MDPI Barcelona, Spain
lucia.russo@mdpi.com
Salehi Rasoul
Isfahan University of Medical Science, Isfahan, Iran
r_salehi@med.mui.ac.ir
Saxena Meera
University of Basel, Basel, Switzerland
meera.saxena@unibas.ch
Schmittgen Thomas
University of Florida, Gainesville, USA
tschmittgen@cop.ufl.edu
Seimiya Makiko
ETH Zurich, Switzerland
makiko.seimiya@bsse.ethz.ch
Shetty Sunil
University of Basel, Switzerland
sunil.shetty@unibas.ch
Skawran Britta
Medical School Hannover, Hannover, Germany
skawran.britta@mh-hannover.de
Slack Frank J.
Harvard Medical School, Boston, USA
fslack@bidmc.harvard.edu
Stiegelbauer Verena
Medical University of Graz, Austria
verena.stiegelbauer@medunigraz.at
Sylvester Bianca
MDPI Romania
bianca.sylvester@mdpi.com
Tahmasebi Maryam
Ahvaz Jundishapur University of Medical Sciences, Ahvaz, Iran
maryam_tahmaseby@yahoo.com
Terreri Sara
Institute of Genetics and Biophysics, Naples, Italy
sara.terreri@igb.cnr.it
Uko Bubaraye
University of Liverpool, Liverpool, UK
b.uko@liv.ac.uk
Vajen Beate
Medical School Hannover, Hannover, Germany
vajen.beate@mh-hannover.de
Vandesompele Jo
Ghent University, Ghent, Belgium
Joke.Vandesompele@UGent.be
Vazquez Franck
MDPI Basel, Switzerland
vazquez@mdpi.com
Veronese Angelo
University of Chieti-Pescara, Italy
a.veronese@unich.it
Wang Yun
Sun Yat-sen University, Guangzhou, China
pemberley05@hotmail.com
Wu Wei-Zhong
Fudan University, Shanghai, China
wu.weizhong@zs-hospital.sh.cn
Wu Zongsong
University of Basel, Switzerland
zongsong.wu@unibas.ch
Yap Yoon Sing
New York University School of Medicine, New York, USA
yoonsing.yap@nyumc.org
Zavolan Mihaela
University of Basel, Switzerland
mihaela.zavolan@unibas.ch
Zeitler Daniela
University of Regensburg, Germany
Daniela.Zeitler@ur.de
Zhang Shu
Xi'an Jiaotong University, China
drzhangshu@163.com
Conference Photos

Welcome Message by Dr. Conradin Cramer, Regierungsrat Kanton Basel-Stadt
Conradin Cramer with George Calin, Chair of ISFMS2017 and John Mattick, Member of the Scientific Committee
Witold Filipowicz, FMI Basel, John Mattick, University of New South Wales and Conradin Cramer
Amy Pasquinelli University of California, San Diego, USA
Coffee Break at the Biocenter, University of Basel
Qihong Huang, The Wistar Institute, Philadelphia, USA
Sven Diederichs, DKFZ, Heidelberg, Germany
Carlo Croce, The Ohio State University, Columbus, USA
Poster Exhibition
The conference team. Lucia Russo, Daniel Luong and Michèle Degen
John Mattick and Franck Vazquez
Helge Grosshans, FMI Basel, Switzerland
Fabbri Muller, University of Southern California, Los Angeles, USA
John Mattick
Franck Slack and Helge Grosshans
Mihaela Zavolan, University of Basel, Switzerland
George Calin, The University of Texas MD Anderson Cancer Center, Houston, TX, USA
Franck Vazquez, Lucia Russo, Michèle Degen and Matthias Burkhalter, MDPI AG, Basel and Barcelona
Tom Schmittgen, University of Florida, Gainesville, USA
Final talks at ISFMS2017
Jo Vandesompele, Ghent University, Belgium
Tom Schmittgen and George Calin
Sara Terreri, Winner of the Best Poster Award
Franck Vazquez, CEO, MDPI AG, Basel, Switzerland
George Calin and Franck Vazquez
Conference Award
The Sponsors, IJMS, Cancers and Non-Coding RNA, offers one award to our participants at the conference:
1. Best Poster Award (1): 1 waived publication in one of our journals. The nominations have been assessed by the Organizing Committee.
Winner: Sara Terreri, Institute of Genetics and Biophysics (IGB-ABT), National Research Council (CNR), Naples, Italy
Sara Terreri the Winner of the 'Best Poster Award' together with George Calin, Chair ISFMS2017 and Franck Vazquez, CEO MDPI AG
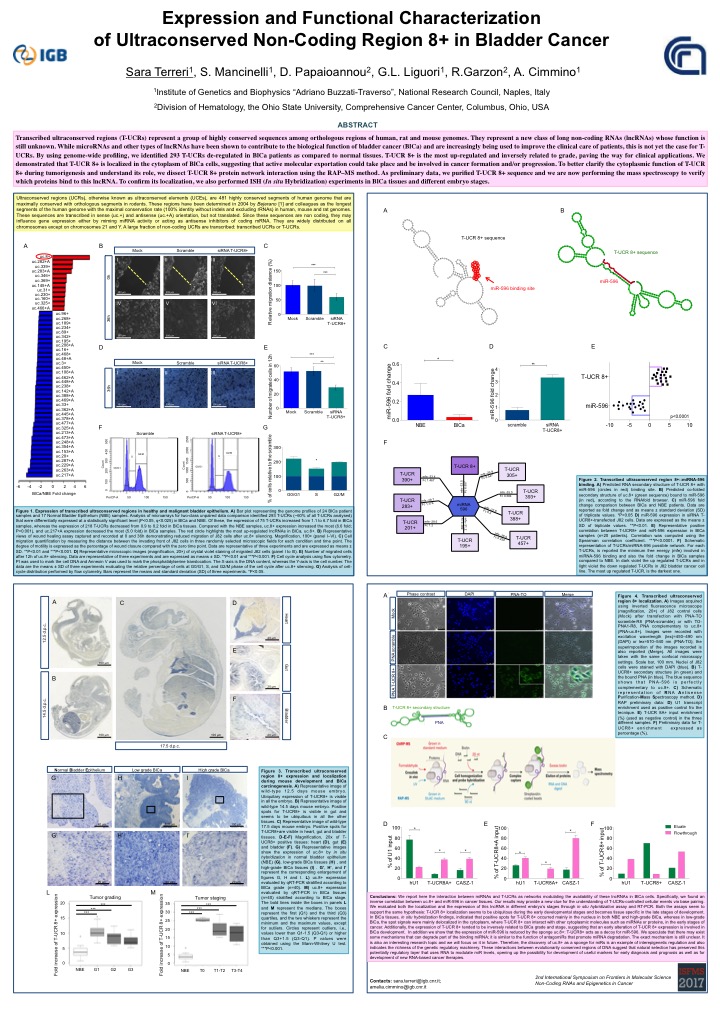
Expression and Functional Characterization of Ultraconserved Non-Coding Regions 8+ in Bladder Cancer
Sara Terreri1, Sara Mancinelli1, Dimitrios Papaioannou2, Giovanna L. Liguori1, Ramiro Garzon2, Amelia Cimmino1
1 Institute of Genetics and Biophysics (IGB-ABT), National Research Council (CNR), Naples, Italy.
2 Division of Hematology, the Ohio State University, Comprehensive Cancer Center, Columbus, Ohio,
USA.
Transcribed ultraconserved regions (T-UCRs) represent a group of highly conserved sequences among orthologous regions of human, rat and mouse genomes. They represent a new class of long non-coding RNAs (lncRNAs) whose function is still unknown. While microRNAs and other types of lncRNAs have been shown to contribute to the biological function of bladder cancer (BlCa) and are increasingly being used to improve the clinical care of patients, this is not yet the case for T-UCRs. By using genome-wide profiling, we identified 293 T-UCRs de-regulated in BlCa patients as compared to normal tissues. T-UCR 8+ is the most up-regulated and inversely related to grade, paving the way for clinical applications. We demonstrated that T-UCR 8+ is localized in the cytoplasm of BlCa cells, suggesting that active molecular exportation could take place and be involved in cancer formation and/or progression. To better clarify the cytoplasmic function of T-UCR 8+ during tumorigenesis and understand its role, we dissect T-UCR 8+ protein network interaction using the RAP–MS method. As preliminary data, we purified T-UCR 8+ sequence and we are now performing the mass spectroscopy to verify which proteins bind to this lncRNA. To confirm its localization, we also performed ISH (In situ Hybridization) experiments in BlCa tissues and different embryo stages.
Conference Video

A conference video will be uploaded soon.
Travel & Registration Information
To register with this conference, please follow this link. If you need assistance, please e-mail the Conference Secretariat.
Early Registration Fees (until 5 May 2017)
- Academic, early: 700.00 CHF
- Scientific Board Members of the journal IJMS, early: 350.00 CHF
- MDPI Author* or Reviewer**, early: 600.00 CHF
- Student, early: 550.00 CHF
- Non-academic, early: 1100.00 CHF
Late Registration Fees (from 6 May 2017)
- Academic, Late: 800.00 CHF
- Scientific Board Members of the journal IJMS, late: 450.00 CHF
- MDPI Author* or Reviewer**, late: 700.00 CHF
- Non-academic, late: 1200.00 CHF
Waived Fees
- Waived Fees - Invited Speakers and MDPI Guests: 0.00 CHF: 0.00 CHF
Sponsored Rate (20 available)
- Special (0/20): 100.00 CHF
Conference Dinner
- Conference Dinner: 80.00 CHF
The registration fee must be paid latest one month after registration and includes attendance of all conference sessions, morning/afternoon coffee breaks and lunches, conference bag and program book. Other dinners are not included and are left at the convenience of attendees
For onsite registration, please contact the Conference Secretariat.
* First and last authors of articles published in 2014 or later will enjoy a discount.
**Reviewers who provided timely review reports in 2014 or later will enjoy a discount.
Prices are in CHF (Swiss Francs)
Cancellation Policy
Participation to the conference is considered final only once the registration fees have been paid. The number of participants is limited, once the number of paid registrations reaches the maximum number of participants, unpaid registrations will be cancelled.
Cancellation of paid registration is possible under the terms listed below:> 2 months before the conferenceFull refund but 100 CHF are retained for administration> 1 month before the conferenceRefund 50% of the applying fees> 2 weeks before the conferenceRefund 25% of the applying fees< 2 weeks before the conferenceNo refund
Visa Requirements
Citizens of the EU and the EFTA and their family members
- Citizens of EU and EFTA countries must present valid personal identification, an ID card or a passport. No visa required.
- Regardless of their citizenship, family members of EU and EFTA citizens must present a valid passport and they must have a specific residence permit from a Schengen country (no visa required). Otherwise a visa must be obtained.
For more information, see https://www.bfm.admin.ch/content/bfm/en/home/themen/einreise/kurzfristig/eu-efta-buerger.html.
- EU Countries: Austria, Belgium, Bulgaria, Croatia, Cyprus, Czech Republic, Denmark, Estonia, Finland, France, Germany, Greece, Hungary, Ireland, Italy, Latvia, Lithuania, Luxembourg, Malta, Netherlands, Poland, Portugal, Romania, Slovakia, Slovenia, Spain, Sweden, United Kingdom.
- EFTA Countries: Iceland, Liechtenstein, Norway, Switzerland.
- Schengen Countries: Austria, Belgium, Czech Republic, Denmark, Estonia, Finland, France, Germany, Greece, Hungary, Iceland, Italy, Latvia, Liechtenstein, Lithuania, Luxembourg, Malta, Netherlands, Norway, Poland, Portugal, Slovakia, Slovenia, Spain, Sweden, Switzerland.
Citizens of the USA and Canada
A valid travel passport is required. No visa required for stays of up to 90 days.
Third-Country Nationals
A valid travel passport is required and in some cases a visa must be obtained prior to entering Switzerland. For more information, check the column "Visa required for stays of up to 90 days" for your country in the Overview of ID and visa provisions according to nationality from the Swiss Office for Migration.
Traveling to Basel
For a complete overview, see wikitravel.org.
- By Plane: Basel can easily be reached by direct flights from many cities within Europe to the EuroAirport Basel-Mulhouse-Freiburg (IATA airport codes: MHL/BSL/EAP). Inter-continental travel to Basel is recommended via Amsterdam, Frankfurt, Paris or London airports. Remember to use the “Swiss Exit”, not the exit to France.
Basel is readily connected within 30 minutes via the airport bus line 50 or by taxi.Another airport in the area includes Zurich International (ZRH).
- By Train: reachable from most major cities in France (via Strasbourg or Mulhouse), Germany (via Frankfurt a. M. or Mannheim) and Italy (via Milan). The two train stations Basel SBB and Basel Bad. are both conveniently connected with the city public transports (BVB). A mobility ticket for free use of public transport is provided by many hotels in Basel.
Conference Organizers
Chairman
Prof. George A. Calin
The University of Texas MD Anderson Cancer Center, Houston, TX, USA
Scientific Committee
Prof. John S. Mattick
Institute Director and Head, RNA Biology and Plasticity Laboratory
Senior Principal Research Fellow, Garvan Institute of Medical Research
University of New South Wales, Sydney, Australia
Prof. Manel Esteller
Leader of the Cancer Epigenetics Group
Bellvitge Biomedical Research Institute (IDIBELL)
Barcelona, Spain
Dr. Massimo Negrini
Department of Morphology
Surgery and Experimental Medicine
University of Ferrara, Italy
Prof. Thomas Schmittgen
Professor Department of Pharmaceutics, College of Pharmacy
University of Florida , USA
Prof. Frank J. Slack
Professor, Pathology, Harvard Medical School
Professor, Pathology, Beth Israel Deaconess Medical Center
Conference Secretariat
Dr. Franck Vazquez
Dr. Lucia Russo
Ms. Rayna Ren
Ms. Michèle Degen
Mr. Matthias Burkhalter
E-mail: ncrna-cancer@mdpi.com
Tel. +41 61 683 77 35
Mailing Address
MDPI AG
ISFMS 2017 Secretariat
St. Alban-Anlage 66
4052 Basel, Switzerland
www.mdpi.com
Conference Schedule
The program of the 2nd International Symposium on Frontiers in Molecular Science 'Non-Coding RNAs and Epigenetics in Cancer', Basel, is now available in the abstract book (last update June, 15). 
Session Start
Wednesday 21 June 2017: 08:30 - 13:20 / 14:20 - 18:10
Thursday 22 June 2017: 09:00 - 13:20 / 14:20 - 18:10
Friday 23 June 2017: 09:00 - 13:20
Detailed Program
Day 1: Wednesday 21 June 2017
07:30–08:30 Check-in
08:30–08:45 Welcome – Conradin Cramer
08:45–09:00 Introduction – George A. Calin
|
09:00–13:20 Session 1: The Biology of ncRNAs – Sponsored by Arraystar |
09:00–09:30 Reuven Agami – Functional Genetic Screens of Regulatory DNA Elements
09:30–10:00 Amy Pasquinelli – Elucidating the Dark Side of the MicroRNA
10:00–10:20 Joost Kluiver – ZDHHC11 and ZDHHC11B are Novel Critical Components of the Oncogenic miR-150-MYBnetwork in Burkitt Lymphoma
10:20–10:40 Eleonora Candi – Ultra-Conserved Non-Coding Transcript T-UC291 Controls Somatic Tissue Differentiation by Interfering with ACTL6A
10:40–11:00 Carmen Jeronimo – A Multiplatform Approach Identifies miR-152 as a Novel Epigenetically Downregulated microRNA in Prostate Cancer
11:00–11:40 Coffee Break
11:40–12:10 Qihong Huang – Lost in Translation: Long Non-coding RNAs in Metastasis
12:10–12:40 Sven Diederichs – MALAT1 and Beyond - Long Non-coding RNAs in Lung Cancer
12:40–13:00 Daniela Zeitler – Hyper-phosphorylation of Argonaute Proteins Affects mRNA Binding and is Essential for microRNA-guided Gene Silencing
13:00–13:20 Jelena Kresoja-Rakic – lncRNA RP11-334E6.12 expression is highly correlated with increased THY-1 expression in chemoresistant primary mesothelioma cells
13:20–14:30 Lunch and Poster Session
14:30–18:20 Session 2: Translational Applications of ncRNAs as Biomarkers
Session Chairs: Carlo Croce and Menashe Bar-Eli
14:30–15:00 Carlo Croce – MicroRNA Dysregulation to Identify Therapeutic Target Combinations for Chronic Lymphocytic Leukemia
15:00–15:30 Menashe Bar-Eli – RNA Editing and Melanoma Metastasis
15:30–15:50 Stefan Eichmüller – miRNAs Modulating Melanoma Cell Invasion
15:50–16:30 Coffee Break
16:30–17:00 Helge Grosshans – Noncoding RNA Function and Regulation in Animal Development
17:00–17:30 Fabbri Muller – Role of Exosomal miRNAs in the Biology of the Tumor Microenvironment
17:30–18:00 Manuela Ferracin – Epigenetic Biomarkers of Prognosis in Stage IIA Colon Cancer
18:00–18:20 Dror Avni – Alterations of MicroRNAs Throughout the Malignant Evolution of Cutaneous Squamous Cell Carcinoma: The Role of miR-497 in Epithelial to Mesenchymal Transition of Keratinocytes
Day 2: Thursday 22 June 2017
09:00–13:10 Session 3: Bioinformatics and ncRNAs World
Session Chairs: Isidore Rigoutsos and John S. Mattick
09:00–09:30 Isidore Rigoutsos – Transcriptomic Heterogeneity: Known and Novel Short Non-coding Regulatory RNAs that Depend on Sex, Population Origin, Tissue, and Disease
09:30–10:00 Mihaela Zavolan – The 3' UTR Landscape of Human Cancers
10:00–10:20 Giovanna Brancati – Fatal Imperfections: Determinants of miRNA Target Specificity
10:20–10:40 Francisco J. Enguita – miRNAtools: Advanced Training Using the miRNA Web of Knowledge
10:40–11:00 Rory Johnson – Cancer Driver Long Noncoding RNA Discovery in the Pan-Cancer Analysis of Whole Genomes (PCAWG) Collaboration
11:00–11:40 Coffee Break
11:40–12:10 John S. Mattick – Exons are the modular unit of structure-function in regulatory RNAs
12:10–12:40 Ulf Andersson Orom – Chromatin-Release Is Important for Long NcRNA Function
12:40–13:10 Manel Esteller – Epigenetics and Epitranscriptomics of Non-Coding RNAs in Human Cancer
13:10–14:10 Lunch and Poster Session
14:10–17:50 Session 4: ncRNA Therapeutics
Session Chairs: Frank J. Slack and Massimo Negrini
14:10–14:40 Frank J. Slack – MicroRNA-Based Therapeutics in Cancer
14:40–15:10 George A. Calin – About Chomsky, Non-Coding RNA Structure and Cancer Patient's Treatment
15:10–15:30 Aniello Russo – Role of miR-125a in Hepatic Carcinogenesis
15:30–16:10 Coffee Break
16:10–16:40 Massimo Negrini – Multiple Approaches for miRNA-based Therapies of Cancer
16:40–17:10 Kalpana Ghoshal – Transcriptome-Wide Mapping of the miR-122 Targetome Revealed its Mechanistic Role in the Maintenance of Liver Homeostasis and Suppressing Hepatocarcnogenesis
17:10–17:30 Silvia Catuogno – Development of RNA Aptamers for Targeting B-cell-derived Malignancies
17:30–17:50 Alva Rani James – A Specific long non-coding RNA Expression Signature Defines the Philadelphia-like B-cell Acute Lymphoblastic Leukemia Subtype
19:00 Conference Dinner
Day 3: Friday 23 June 2017
09:00–13:20 Session 5: ncRNA Technologies
Session Chairs: Thomas Schmittgen and Manel Esteller
09:00–09:30 Thomas Schmittgen – microRNAs Shape Plasticity of Pancreatic Acini
09:30–10:00 Jo Vandesompele – Tools for lncRNA Research in Cancer
10:00–10:20 Britta Skawran – The microRNA-449 Family Inhibits TGF-β-mediated Liver Cancer Cell Migration by Targeting SOX4
10:20–10:40 Jörg Krummheuer – An Improved Algorithm for Antisense LNA™ GapmeR Design
10:40–11:00 Mattia Boeri – High Risk Plasma microRNA Signature is Associated with an Immune-related Gene Expression Profile of Lung Tumour Tissues
11:00–11:40 Coffee Break
11:40–12:00 Martin Pichler – MicroRNA-196 Influence Metastases Formation in Colorectal Cancer through Regulation of HOXB and GALNT Gene Expression
12:00–12:30 Maite Huarte Martinez – Functional long Noncoding RNAs in Cancer Pathways
12:30–12.50 Enrique Fuentes-Mattei – Plasma Viral miRNAs as Targeted-biomarkers of Occult Viral Infections Prevalence and Sepsis Aggressiveness
12:50–13:10 Paola Parrella – Stepwise Analysis of MIR9 Loci Identifies miR-9-5p to be Involved in Oestrogen Regulated Pathways in Breast Cancer Patients
13:10 Concluding Remarks
Conference Speakers
Session 1: The Biology of ncRNAs - Sponsored by Arraystar
Prof. Reuven Agami - Functional Genetic Screens of Regulatory DNA Elements
Netherlands Cancer Institute - NKI Amsterdam, The Netherlands
Prof. Amy Pasquinelli - Elucidating the Dark Side of the MicroRNA
Division of Biological Sciences, University of California, USA
Prof. Qihong Huang - Lost in Translation: Long Non-Coding RNAs in Metastasis
Tumor Microenvironment and Metastasis Program, The Wistar Institute, Philadelphia, USA
Prof. Sven Diederichs - MALAT1 and Beyond - Long Non-Coding RNAs in Lung Cancer
Division Head RNA Biology & Cancer, DKFZ - Deutsches Krebsforschungszentrum, Germany
Session 2: Translational Applications of ncRNAs as Biomarkers
Prof. Carlo Croce - MicroRNA Dysregulation to Identify Therapeutic Target Combinations for Chronic Lymphocytic Leukemia
Department of Cancer Biology and Genetics, The Ohio State University, USA
Prof. Menashe Bar-Eli - RNA Editing and Melanoma Metastasis
Department of Cancer Biology, The University of Texas, MD Anderson Cancer Center, USA
Prof. Helge Grosshans - Non-Coding RNA Function and Regulation in Animal Development
Friedrich Miescher Institute for Biomedical Research (FMI) Basel, Switzerland
Prof. Fabbri Muller - Role of Exosomal miRNAs in the Biology of the Tumor Microenvironment
Pediatrics and Molecular Microbiology and Immunology, Children's Hospital Los Angeles, University of Southern California, USA
Prof. Manuela Ferracin - Epigenetic Biomarkers of Prognosis in Stage IIA Colon Cancer
Department of Specialised, Experimental, and Diagnostic Medicine, University of Bologna, Italy
Session 3: Bioinformatics of ncRNAs
Prof. Isidore Rigoutsos - Transcriptomic Heterogeneity: Known and Novel Short Non-Coding Regulatory RNAs that Depend on Sex, Population Origin, Tissue, and Disease
Director Computational Medicine Center, Thomas Jefferson University
Prof. Mihaela Zavolan - The 3' UTR Landscape of Human Cancers
Professor in Computational Biology/Genomics, Biozentrum, University of Basel, Switzerland
Prof. John S. Mattick - Exons are the Modular Unit of Structure-Function in Regulatory RNAs
Institute Director and Head, RNA Biology and Plasticity Laboratory Senior Principal Research Fellow, Garvan Institute of Medical Research, University of New South Wales, Sydney, Australia
Prof. Ulf Andersson Orom - Chromatin-Release Is Important for Long ncRNA Function
Long non-coding RNA Research Group, Max Planck Institute for Molecular Genetics, Germany
Prof. Manel Esteller - Epigenetics and Epitranscriptomics of Non-Coding RNAs in Human Cancer
Leader of the Cancer Epigenetics Group, Bellvitge Biomedical Research Institute (IDIBELL), Barcelona, Spain
Session 4: ncRNA Therapeutics
Prof. Frank J. Slack - MicroRNA-Based, Therapeutics in Cancer
Professor, Pathology, Harvard Medical School, Professor, Pathology, Beth Israel Deaconess Medical Center, Boston, USA
Prof. George A. Calin - About Chomsky, Non-Coding RNA Structure and Cancer Patient's Treatment
The University of Texas, MD Anderson Cancer Center, Houston, TX, USA
Prof. Massimo Negrini - Multiple Approaches for miRNA-based Therapies of Cancer
Department of Morphology Surgery and Experimental Medicine, University of Ferrara, Italy
Prof. Kalpana Ghoshal - Transcriptome-Wide Mapping of the miR-122 Targetome Revealed Its Mechanistic Role in the Maintenance of Liver Homeostasis and Suppressing Hepatocarcnogenesis
Department of Pathology, The Ohio State University, USA
Session 5: ncRNA Technologies
Prof. Thomas Schmittgen - MicroRNAs Shape Plasticity of Pancreatic Acini
College of Pharmacy, University of Florida, Gainesville, USA
Prof. Jo Vandesompele - Tools for lncRNA Research in Cancer
Chair of the Cancer Research Institute, Ghent University, Belgium
Prof. Maite Huarte Martinez - Functional long Non-Coding RNAs in Cancer Pathways
Center for Applied Medical Research (CIMA), University of Navarra, Spain
P. Poster Exhibition
Show all accepted abstracts (19) Hide accepted abstracts (19)
List of Accepted Abstracts (19) Toggle list
S1. The Biology of ncRNAs - Sponsored by Arraystar Inc.
Show all accepted abstracts (10) Hide accepted abstracts (10)
List of Accepted Abstracts (10) Toggle list
S2. Translational Applications of ncRNAs as Biomarkers
Show all accepted abstracts (8) Hide accepted abstracts (8)
List of Accepted Abstracts (8) Toggle list
S3. Bioinformatics of ncRNAs
Show all accepted abstracts (7) Hide accepted abstracts (7)
List of Accepted Abstracts (7) Toggle list
S4. ncRNA Therapeutics
Show all accepted abstracts (7) Hide accepted abstracts (7)
List of Accepted Abstracts (7) Toggle list
S5. ncRNA Technologies
Show all accepted abstracts (10) Hide accepted abstracts (10)
List of Accepted Abstracts (10) Toggle list

















